Main Points:
- Old human serum can induce multiple hallmarks of aging in human tissues within days
- Adipose tissue acts as an upstream amplifier of systemic aging signals
- Aging can propagate between tissues through inflammatory and metabolic signaling
- Targeting senescence pathways like CDKN2A may partially reverse these effects
- Organ-chip systems may redefine how aging therapies are developed and tested
The body ages all at once. Or does it?
Aging has long been treated as a slow accumulation of damage, a quiet drift measured in decades. But what if aging could be imposed, rapidly, almost on demand?
In a recent study published in Nature Biomedical Engineering, researchers built a human tissue system that appears to do exactly that. By exposing engineered human tissues to the blood serum of older individuals, they induced a cascade of aging features in just four days. Cells began to behave as though they had lived decades longer than they had.
This isn’t just a technical curiosity. It challenges the implicit timeline of aging itself and reframes systemic aging as something actively maintained, not merely accumulated.
Aging, in this view, is not only what happens within cells. It is also what happens between them.
Aging as a circulating signal, not just a cellular fate
To understand the implications, it helps to revisit what aging is thought to be. Modern geroscience frames aging through a set of biological hallmarks, including cellular senescence, where damaged cells stop dividing but remain metabolically active, and chronic inflammation, where immune signaling becomes persistently elevated. Other hallmarks include metabolic dysfunction, genomic instability, and altered intercellular communication.
That last category, altered communication, has historically been harder to pin down. It implies that aging is not just intrinsic to cells but emerges from signals exchanged across tissues. Blood, as the body’s communication medium, becomes a prime suspect.
Earlier experiments in mice hinted at this. When the circulatory systems of young and old mice were connected, a procedure known as parabiosis, older tissues showed signs of rejuvenation. But translation to humans has remained speculative, limited by the inability to directly test such interventions.
This is the gap the new study attempts to close.
Building aging in a dish, then accelerating it
The researchers constructed a microphysiological system, often referred to as an organ chip, using human induced pluripotent stem cells. These stem cells were differentiated into two key tissues: white adipose tissue, the body’s primary fat storage organ, and liver tissue, a central hub for metabolism.
White adipose tissue is not just a passive fat depot. It actively secretes hormones and inflammatory signals, making it a major regulator of systemic metabolism. The liver, meanwhile, controls glucose production and lipid processing, making it highly sensitive to systemic changes.
The experimental design was deceptively simple. These tissues were exposed to media containing 5% human serum, either from young donors aged 21–34 or older donors over 62. After just four days, the researchers began measuring markers of aging.
They didn’t look for one signal. They looked for many, across multiple hallmarks.
When young tissue starts behaving old
The results were not subtle.
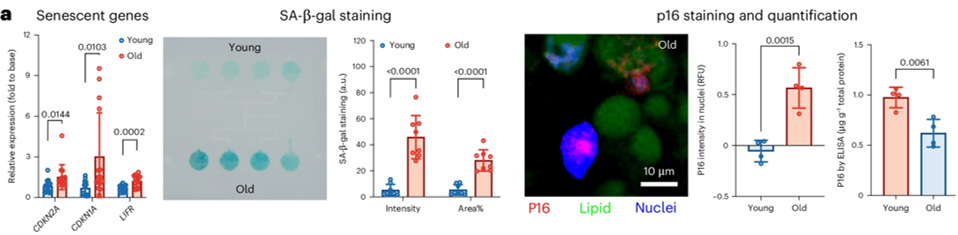
Cells exposed to older serum showed increased levels of SA-β-gal, a commonly used marker of cellular senescence indicating lysosomal activity in non-dividing cells. They also upregulated CDKN2A, a gene encoding the protein p16, which enforces cell cycle arrest, and CDKN1A, which encodes p21, another inhibitor of cell division.
Inflammatory signals surged. IL-6, a cytokine that promotes chronic inflammation, and TNF, a key immune regulator, both increased. These are canonical components of the senescence-associated secretory phenotype (SASP), a pro-inflammatory state that senescent cells adopt and broadcast to neighboring cells.
At the level of DNA, oxidative damage rose. The molecule 8-OHdG, a marker of oxidative lesions in DNA, increased significantly in adipose tissue. This suggests that the serum environment itself is capable of inducing genomic stress.
Metabolism shifted as well. Genes like PPARG, a master regulator of fat cell differentiation, and FABP4, which binds fatty acids inside cells, were upregulated. Lipid uptake increased, and glucose handling deteriorated. The liver tissue began producing more glucose via PCK1, a key enzyme in gluconeogenesis, indicating insulin resistance.
These are not isolated features. They are coordinated hallmarks of aging.
The surprising hierarchy: fat leads, liver follows
One of the study’s most consequential findings emerged when the researchers connected the two tissues.
Adipose tissue exposed to old serum began secreting factors that, when transferred to liver tissue, induced aging-like changes there as well. Liver cells showed increased IL-6, CDKN2A, and SREBP1c, a transcription factor that drives lipid synthesis. They accumulated fat and became less responsive to insulin.
This suggests a directional model of systemic aging:
Old serum → adipose dysfunction → liver dysfunction
Adipose tissue, often dismissed as excess storage, appears to act as an upstream amplifier. It translates systemic signals into local dysfunction and then propagates those signals outward.
This aligns with epidemiological observations linking visceral fat to metabolic disease. But here, the causality is experimentally reconstructed.
Measuring age not in years, but in expression patterns
To validate whether these changes truly resembled aging, the researchers turned to transcriptomics, the study of gene expression.
They compared RNA profiles from their organ-chip tissues to human data from the GTEx project, a large database of gene expression across human tissues. The overlap was striking. Aging-associated pathways, including TNF signaling, TGFβ signaling, and cellular senescence, were enriched in both datasets.
They then trained a machine learning model to predict biological age based on gene expression. This model achieved up to 97% accuracy in distinguishing young from old human adipose samples.
When applied to the organ-chip tissues, the model showed that tissues exposed to old serum rapidly shifted toward an older predicted age. Conversely, removing old serum or replacing it with young serum partially reversed this shift.
Age, at least at the molecular level, appears more plastic than previously assumed.
Intervening in the signal, not just the cell
The study also tested interventions.
They used dasatinib and quercetin, a senolytic combination known to selectively eliminate senescent cells, as well as rapamycin, which inhibits mTOR signaling, a central regulator of growth and metabolism. They also tested oxytocin, a hormone better known for its role in social bonding but increasingly studied for metabolic effects.
Oxytocin showed the most pronounced improvements in this system, reducing senescence markers and improving metabolic function under certain conditions. Additionally, knocking down CDKN2A reduced aging phenotypes, particularly when old serum was removed.
These interventions do not “reverse aging” in a clinical sense. But they demonstrate that the system is manipulable, and that targeting specific nodes can alter the trajectory of aging signals.
What this changes about systemic aging
The central shift is conceptual.
Aging is often framed as a cell-autonomous process, driven by internal damage. This study supports a complementary model in which aging is also systemically imposed, through circulating factors that coordinate dysfunction across tissues.
This doesn’t negate intracellular damage. It reframes it within a broader network.
It also raises the possibility that interventions could target the environment of cells, not just the cells themselves. Removing pro-aging signals, diluting them, or blocking their downstream effects could be as important as repairing damage.
Where the model falls short
Despite its strengths, the system is still a simplification.
The organ-chip lacks immune cells, vascular structures, and neural inputs, all of which play critical roles in aging. The adipose tissue and liver are composed of iPSC-derived cells, which may not fully recapitulate mature human tissue behavior.
The study also relies on bulk RNA sequencing, which averages signals across cell populations. This obscures cell-type-specific responses, which are increasingly recognized as critical in aging biology.
And while serum from older donors induces aging-like changes, the specific molecules responsible remain unidentified. The signal is real, but its composition is still unresolved.
What this means for longevity science
The implications are less about immediate therapies and more about trajectory.
If systemic aging can be modeled in human tissue within days, it becomes possible to test interventions rapidly and iteratively. This could compress the development timeline for geroprotective drugs.
It also elevates adipose tissue as a central player in aging, not just in obesity or metabolic disease, but in systemic decline.
Most importantly, it suggests that aging is not only a story of accumulation, but also of communication. Cells are not aging in isolation. They are responding to signals, broadcasting them, and amplifying them.
Understanding those signals may be one of the most direct paths to intervening in aging itself.
References
- Qi, L., He, Y., Sviercovich, A., Mei, X., Chen, E., Xia, Y., Conboy, M. J., Conboy, I. M., & Stahl, A. (2026). Human microphysiological systems of aging recreate the in vivo process expediting evaluation of anti-geronic strategies. Nature Biomedical Engineering.
- López-Otín, C., Blasco, M. A., Partridge, L., Serrano, M., & Kroemer, G. (2023). Hallmarks of aging: An expanding universe. Cell, 186(2), 243–278.
- López-Otín, C., Blasco, M. A., Partridge, L., Serrano, M., & Kroemer, G. (2013). The hallmarks of aging. Cell, 153(6), 1194–1217.
- Conboy, I. M., Conboy, M. J., Wagers, A. J., Girma, E. R., Weissman, I. L., & Rando, T. A. (2005). Rejuvenation of aged progenitor cells by exposure to a young systemic environment. Nature, 433(7027), 760–764
- Pálovics, R., Keller, A., Schaum, N., Tan, W., Fehlmann, T., Borja, M., Kern, F., Bonanno, L., Calcuttawala, K., Webber, J., McGeever, A., Luo, J., Pisco, A. O., Karkanias, J., Neff, N. F., Darmanis, S., Quake, S. R., & Wyss-Coray, T. (2022). Molecular hallmarks of heterochronic parabiosis at single-cell resolution. Nature, 603(7900), 309–314.
- The GTEx Consortium. (2020). The GTEx Consortium atlas of genetic regulatory effects across human tissues. Science, 369(6509), 1318–1330.
- Xu, M., Pirtskhalava, T., Farr, J. N., Weigand, B. M., Palmer, A. K., Weivoda, M. M., Inman, C. L., Ogrodnik, M. B., Hachfeld, C. M., Fraser, D. G., Onken, J. L., Johnson, K. O., Verzosa, G. C., Langhi, L. G. P., Weigl, M., Giorgadze, N., LeBrasseur, N. K., Miller, J. D., Jurk, D., … Kirkland, J. L. (2018). Senolytics improve physical function and increase lifespan in old age. Nature Medicine, 24(8), 1246–1256.